Introduction: what are protein domains?
Protein domains are structural units of proteins that can fold, function and evolve independently. [1] Different domains have a particular function, which contributes to the overall role of a protein. [2] Many domains are not unique to one protein, but can appear in many different proteins. The domain names usually derive from their primary biological function in the protein. [3]
Domains of HTRA1 and their functions
IGFBP_N_2 :These domains are insulin growth factor-binding protein homologues that has high affinity binding partners of insulin-like growth factors. As the name suggests, it binds to insulin-like growth factors. The IGFBP N-terminal domain mediates interaction with TSC2 substrate.
KAZAL_2: KAZAL domains are Kazal type serine protease inhibitors with protein binding function. They interact from the canonical proteinase-inhibitor binding loop, where P1 residue has a predominant role (the residue at the P1 position contributing the carbonyl portion to the reactive-site peptide bond). This domain involves in various physiological pathways including complement and coagulation cascades, ECM-receptor interaction and TGF-beta signaling pathway. [4]
Trypsin_2: Trypsin-like peptidase domain that acts as serine proteases. The catalytic activity of the serine proteases from the trypsin family is provided by a charge relay system involving an aspartic acid residue hydrogen-bonded to a histidine, which itself is hydrogen-bonded to a serine. In the ileum, it may be involved in defensin processing, including DEFA5. [5]
PDZ: PDZ domains (also known as Discs-large homologous regions (DHR) or GLGF)) are found in diverse signaling proteins in bacteria, yeasts, plants, insects and vertebrates with protein binding function. They are frequently associated with the plasma membrane. Some PDZs have been shown to bind C-terminal polypeptides; others appear to bind internal (non-C-terminal) polypeptides. Different PDZs possess different binding specificities. [6]
KAZAL_2: KAZAL domains are Kazal type serine protease inhibitors with protein binding function. They interact from the canonical proteinase-inhibitor binding loop, where P1 residue has a predominant role (the residue at the P1 position contributing the carbonyl portion to the reactive-site peptide bond). This domain involves in various physiological pathways including complement and coagulation cascades, ECM-receptor interaction and TGF-beta signaling pathway. [4]
Trypsin_2: Trypsin-like peptidase domain that acts as serine proteases. The catalytic activity of the serine proteases from the trypsin family is provided by a charge relay system involving an aspartic acid residue hydrogen-bonded to a histidine, which itself is hydrogen-bonded to a serine. In the ileum, it may be involved in defensin processing, including DEFA5. [5]
PDZ: PDZ domains (also known as Discs-large homologous regions (DHR) or GLGF)) are found in diverse signaling proteins in bacteria, yeasts, plants, insects and vertebrates with protein binding function. They are frequently associated with the plasma membrane. Some PDZs have been shown to bind C-terminal polypeptides; others appear to bind internal (non-C-terminal) polypeptides. Different PDZs possess different binding specificities. [6]
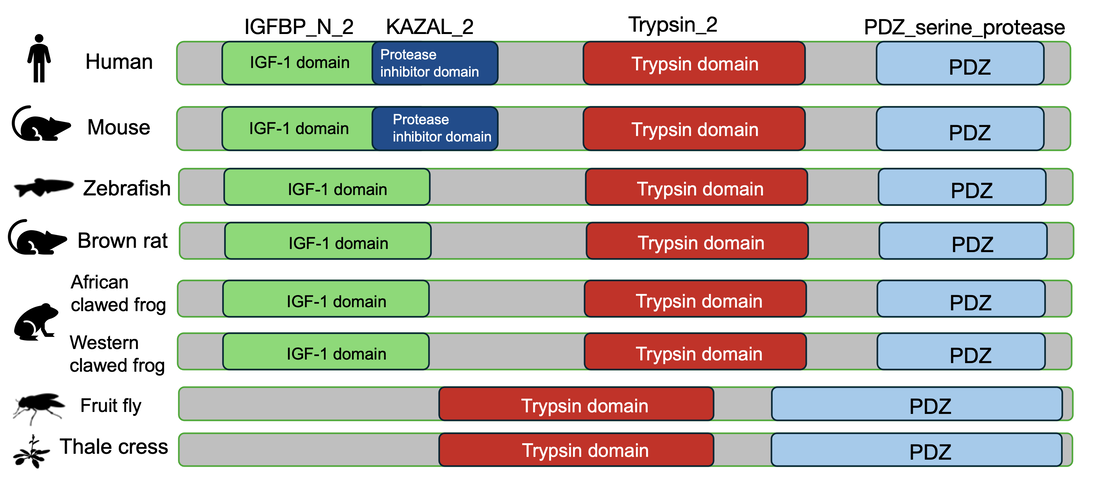 The HTRA1 proteins across different species
The HTRA1 proteins across different species
Discussion
While the Trypsin and PDZ domains of HTRA1 protein are conserved across all model organisms above, IGF-1 and KAZAL domains exist only in certain species. Mice share the closest protein domain structures with human. As species are more phylogenetically similar to human, their protein domains exhibit greater similarity to those found in humans. The observation that certain protein domains in the HTRA1 protein are present in species more closely related to humans (such as mice) but absent in more distant species (like fruit flies) indicates an important aspect of evolutionary conservation. Therefore, IGF-1 and KAZAL domains may evolve uniquely for the survival and adaptation of more complicated organisms.
Reference:
Header: https://www.peakpx.com/en/search?q=protein
[1]Wang, Y., Zhang, H., Zhong, H., & Xue, Z. (2021). Protein domain identification methods and online resources. Computational and Structural Biotechnology Journal, 19, 1145–1153. https://doi.org/10.1016/j.csbj.2021.01.041
[2]EMBL-EBI. (2024). What are protein domains? | Protein classification. Retrieved March 29, 2024, from Ebi.ac.uk website: https://www.ebi.ac.uk/training/online/courses/protein-classification-intro-ebi-resources/protein-classification/what-are-protein-domains/
[3]3.6: Protein Domains, Motifs, and Folds in Protein Structure. (2018, December 20). Retrieved March 29, 2024, from Biology LibreTexts website: https://bio.libretexts.org/Bookshelves/Cell_and_Molecular_Biology/Book%3A_Basic_Cell_and_Molecular_Biology_(Bergtrom)/03%3A_Details_of_Protein_Structure/3.06%3A_Protein_Domains_Motifs_and_Folds_in_Protein_Structure#:~:text=Protein%20motifs%20are%20small%20regions,unique%20chemical%20or%20biological%20function.
[4]SMART: KAZAL domain annotation. (2017). Retrieved March 29, 2024, from Embl-heidelberg.de website: http://smart.embl-heidelberg.de/smart/do_annotation.pl?ACC=SM000280&BLAST=DUMMY
[5]InterPro. (2024). Retrieved March 29, 2024, from Ebi.ac.uk website: https://www.ebi.ac.uk/interpro/protein/UniProt/P07478/
[6]SMART: Error. (2017). Retrieved March 29, 2024, from Embl-heidelberg.de website: http://smart.embl-heidelberg.de/smart/show_motifs.pl
Header: https://www.peakpx.com/en/search?q=protein
[1]Wang, Y., Zhang, H., Zhong, H., & Xue, Z. (2021). Protein domain identification methods and online resources. Computational and Structural Biotechnology Journal, 19, 1145–1153. https://doi.org/10.1016/j.csbj.2021.01.041
[2]EMBL-EBI. (2024). What are protein domains? | Protein classification. Retrieved March 29, 2024, from Ebi.ac.uk website: https://www.ebi.ac.uk/training/online/courses/protein-classification-intro-ebi-resources/protein-classification/what-are-protein-domains/
[3]3.6: Protein Domains, Motifs, and Folds in Protein Structure. (2018, December 20). Retrieved March 29, 2024, from Biology LibreTexts website: https://bio.libretexts.org/Bookshelves/Cell_and_Molecular_Biology/Book%3A_Basic_Cell_and_Molecular_Biology_(Bergtrom)/03%3A_Details_of_Protein_Structure/3.06%3A_Protein_Domains_Motifs_and_Folds_in_Protein_Structure#:~:text=Protein%20motifs%20are%20small%20regions,unique%20chemical%20or%20biological%20function.
[4]SMART: KAZAL domain annotation. (2017). Retrieved March 29, 2024, from Embl-heidelberg.de website: http://smart.embl-heidelberg.de/smart/do_annotation.pl?ACC=SM000280&BLAST=DUMMY
[5]InterPro. (2024). Retrieved March 29, 2024, from Ebi.ac.uk website: https://www.ebi.ac.uk/interpro/protein/UniProt/P07478/
[6]SMART: Error. (2017). Retrieved March 29, 2024, from Embl-heidelberg.de website: http://smart.embl-heidelberg.de/smart/show_motifs.pl
