Introduction: What is Phylogeny?
Phylogeny, or phylogenetic tree, is a studying using a branching diagram that demonstrates the evolutionary history and relationships among different species. As shown in Figure 1, the time goes from left (past) to right (present). A root is where all organisms originate. A node is a branching point from a common ancestor population. A clade is a group of organisms sharing a common ancestor. A taxon is a unit of classified group. There are also different ways to draw phylogenetic trees, as shown in figure 2. [2]
Build phylogenic tree using HTRA1 protein sequence
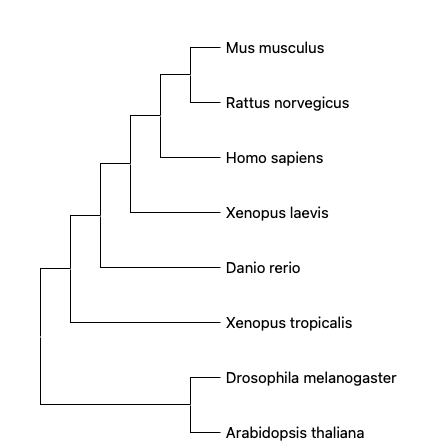
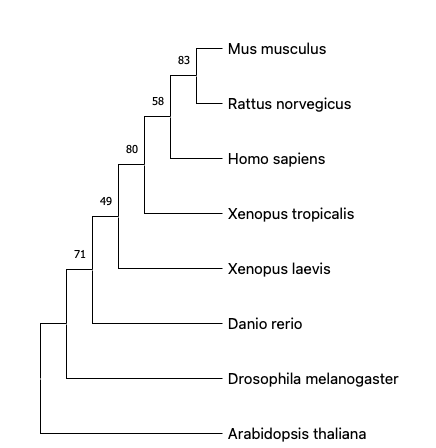
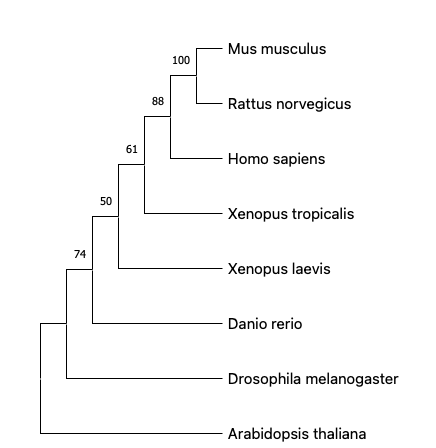
To generate the phylogenic tree, the protein sequences are obtained from FASTA on NCBI and then aligned in a software called MEGA(https://www.megasoftware.net/). The sequence can be aligned using either ClustalW or MUSCLE multiple sequence alignment. MUSCLE uses a progressive algorithm that performs faster than ClustalW and achieves average accuracy similar to ClustalW.[3] The aligned sequence are then analyzed and built into phylogenetic trees using either Neighbor joining or Maximum likelihood phylogenetic inference method. Here are four phylogenetic trees using each alignment and inference method. The numbers above the branches are bootstrap support values.
Discussion
From the phylogenetic trees built using different alignments and inference methods, we can see that HTRA1 is widely conserved across species. Among all the model organisms provided above, mouse and rat share the most recent common ancestor to human, which corresponds with their high identity percentage (90%+) in the homology page and similar protein domains with human HTRA1. This suggests mouse to be one of the model organisms for the study of HTRA1 gene.
Reference:
Header: https://www.earlham.ac.uk/articles/plotting-phylogenetic-trees-r-alternating-clade-highlights
[1]:Bruslind, L. (2019, August). Taxonomy & Evolution. Retrieved March 29, 2024, from Oregonstate.education website: https://open.oregonstate.education/generalmicrobiology/chapter/taxonomy-evolution/
[2]Phylogenetic Trees and Monophyletic Groups | Learn Science at Scitable. (2014). Retrieved March 29, 2024, from Nature.com website: https://www.nature.com/scitable/topicpage/reading-a-phylogenetic-tree-the-meaning-of-41956/
[3]Edgar, R. C. (2004). MUSCLE: a multiple sequence alignment method with reduced time and space complexity. BMC Bioinformatics, 5(1). https://doi.org/10.1186/1471-2105-5-113
Header: https://www.earlham.ac.uk/articles/plotting-phylogenetic-trees-r-alternating-clade-highlights
[1]:Bruslind, L. (2019, August). Taxonomy & Evolution. Retrieved March 29, 2024, from Oregonstate.education website: https://open.oregonstate.education/generalmicrobiology/chapter/taxonomy-evolution/
[2]Phylogenetic Trees and Monophyletic Groups | Learn Science at Scitable. (2014). Retrieved March 29, 2024, from Nature.com website: https://www.nature.com/scitable/topicpage/reading-a-phylogenetic-tree-the-meaning-of-41956/
[3]Edgar, R. C. (2004). MUSCLE: a multiple sequence alignment method with reduced time and space complexity. BMC Bioinformatics, 5(1). https://doi.org/10.1186/1471-2105-5-113